Mini Virtual IFN Immune Cell
Published:
Mini Virtual IFN Immune Cell
Mini Virtual IFN Immune Cell is a perturbation-aware single-cell virtual cell prototype that learns interferon and COVID immune-state transitions and exposes a lightweight inference API for real-cell prediction.
The current v1 setup combines clean interferon stimulation supervision from Kang with heterogeneous immune-state signals from a COVID immune atlas. The goal is to create a compact, biologically interpretable model that can predict response shifts across immune perturbation states without requiring a large whole-cell simulator.
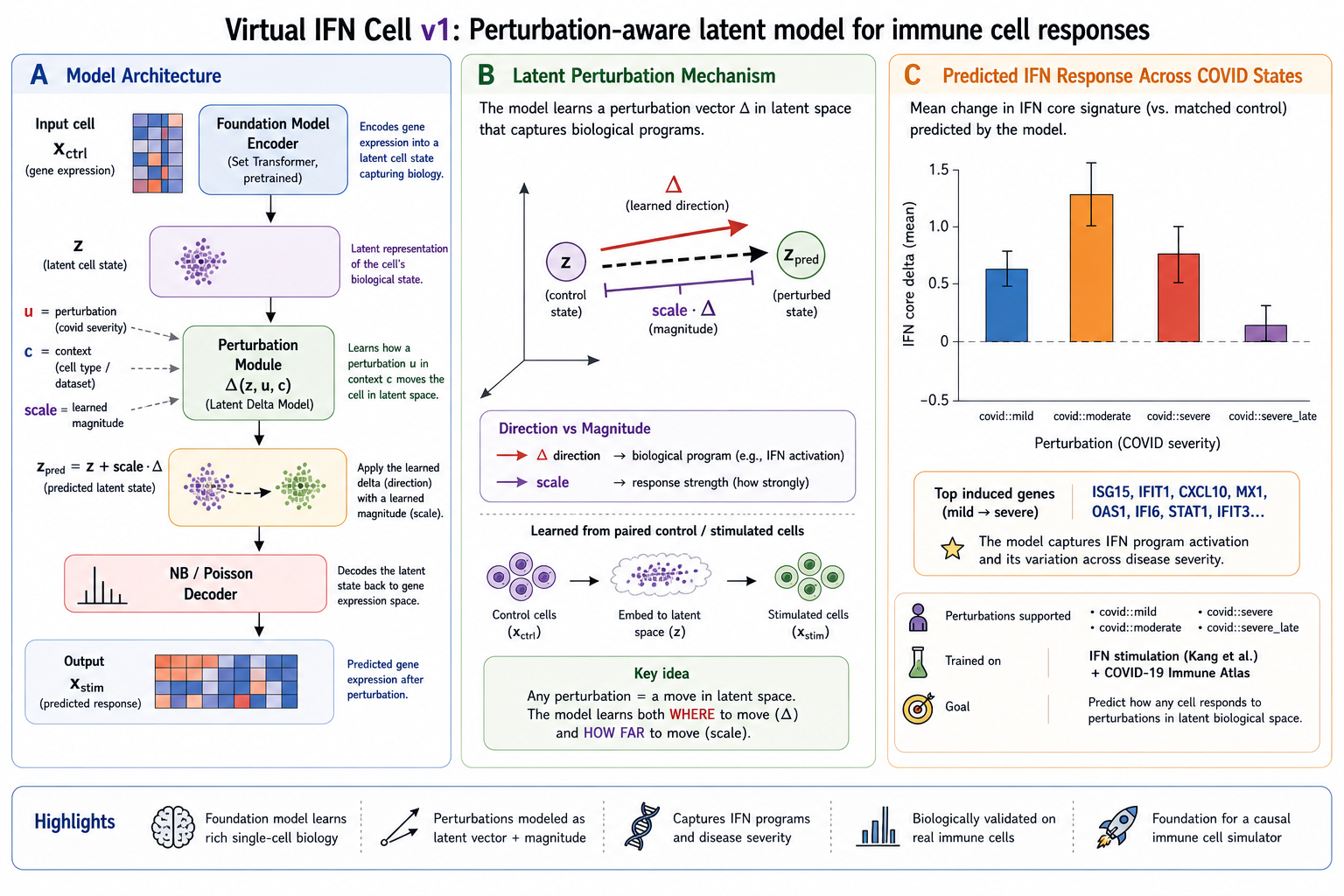
Overview
The current virtual response prediction setup supports COVID perturbation states including:
covid::mildcovid::moderatecovid::severecovid::severe_late
This makes the project useful for exploring how interferon-linked immune programs shift across disease severity, while keeping the representation simple enough to inspect and extend.
What This Project Demonstrates
- latent perturbation modeling for single-cell response prediction
- a real perturbation vocabulary instead of one generic stimulation token
- learned perturbation magnitude calibration
- pathway-level biological validation
- a lightweight real-cell inference API
Why It Matters
Interferon signaling is central to antiviral immunity, inflammatory regulation, and disease progression. In practice, it is often studied either with highly simplified pathway cartoons or with large models that are hard to interpret and hard to deploy.
Mini Virtual IFN Immune Cell aims for a practical middle ground:
- compact enough for rapid experimentation
- structured enough to encode meaningful perturbation states
- interpretable enough for pathway-level biological validation
- flexible enough to support real-cell inference demos
Example Workflow
Observed Cell State
↓
Perturbation Encoder
↓
Latent Response Shift
↓
Predicted Immune State / Gene Program Output
This framing makes it possible to ask practical questions such as:
- How does a real immune cell respond under different COVID-linked perturbation states?
- Does the model capture directionally meaningful IFN response changes?
- Are the learned perturbation magnitudes biologically plausible?
- Can pathway-level validation support qualitative use of the predictions?
v1 Results
The current v1 model is directionally useful and biologically interpretable, but still conservative and imperfect.
- stable delta correlation is roughly in the
0.65-0.67range - IFN pathway correlation is strong enough to support qualitative demos
- predicted IFN response is directionally meaningful across COVID states
- learned scale tends to stay conservative rather than aggressively amplifying responses
- STAT1 knockout behavior is still experimental and weak
Quickstart
Run the real-cell demo after placing the required COVID .h5ad file in the repository root:
python demos/demo_real_covid_cell_api.py
If COVID-19_Immune_Altas.h5ad is not present in the repository root, the demo stops with a clear message.
The COVID atlas used in the current setup was downloaded from Cellxgene: COVID immune atlas collection
Generate Figures
python scripts/make_v1_figures.py
Outputs
The outputs/v1/ folder includes evaluation artifacts for the current model version:
heldout_covid_eval_metrics_seed3.jsonbiological_validation_metrics_seed3.jsondemo_real_cell_prediction.jsondemo_severity_progression.csv
These outputs support both held-out evaluation and qualitative demo workflows.
Training
The current v1 training command is captured in:
configs/v1/train_v1_seed3_epoch10.sh
The training backend used to generate the checkpoint is:
scripts/train_virtual_ifn_cell.py
Design Principles
- Perturbation-aware: learn explicit immune-state transitions rather than a generic stimulation effect
- Biology-grounded: validate predictions at the pathway level, not only through generic metrics
- Lightweight: expose a simple API for real-cell inference and demos
- Extendable: support future expansion to richer pathway supervision and stronger causal simulations
Limitations
- this is not a clinical model
- this is not a validated causal simulator
- STAT1 knockout behavior is exploratory
- severity ordering is not guaranteed to be monotonic
v1is trained on a limited dataset setup and should be interpreted as a prototype
Roadmap
v2: pathway-weighted lossv2: explicit IFN/JAK/STAT perturbation datav2: improved causal gene knockout simulationv2: demo notebook
Data Note
Users should download datasets separately and respect the licenses, terms, and citations of the original data sources. Large .h5ad files are intentionally excluded from the repository.
👉 GitHub Repository: virtual-ifn-cell