MultiOmicsBind: Deep Learning framework for Multi-Omics Data Integration
Published:
MultiOmicsBind: Deep Learning framework for Multi-Omics Data Integration 🧬
Bringing ImageBind’s revolutionary binding modality concept to multi-omics data integration
The Multi-Omics Challenge 🎯 In today’s biological research landscape, we’re drowning in data but thirsting for insights. Researchers routinely generate:
🧪 Genomics data: DNA sequences and variants
📊 Transcriptomics data: Gene expression profiles
🔬 Proteomics data: Protein abundance measurements
⚗️ Metabolomics data: Small molecule concentrations
🖼️ Cell imaging data: Morphological features
The challenge? Each data type tells only part of the biological story. The magic happens when we integrate them all! ✨
Introducing MultiOmicsBind 🚀
MultiOmicsBind is a cutting-edge deep learning framework designed to tackle the multi-omics integration challenge head-on. Built with PyTorch and inspired by ImageBind and the latest advances in contrastive learning, it provides researchers with a powerful, flexible tool for multi-modal biological data analysis.
🌟 Key Features
🔧 Flexible Architecture
- Supports any combination of omics modalities
- Handles varying sample sizes and feature dimensions
- Easily configurable for different research needs
🧠 Advanced Learning
- Contrastive learning for cross-modal representations
- Optional supervised learning for classification tasks
- Robust training with early stopping and scheduling
⚡ User-Friendly
- Comprehensive documentation
- Ready-to-use examples
- Seamless integration with existing workflows
📈 Scalable
- From pilot studies to large biobanks
- Efficient GPU utilization
- Memory-optimized data loading
How It Works 🔍
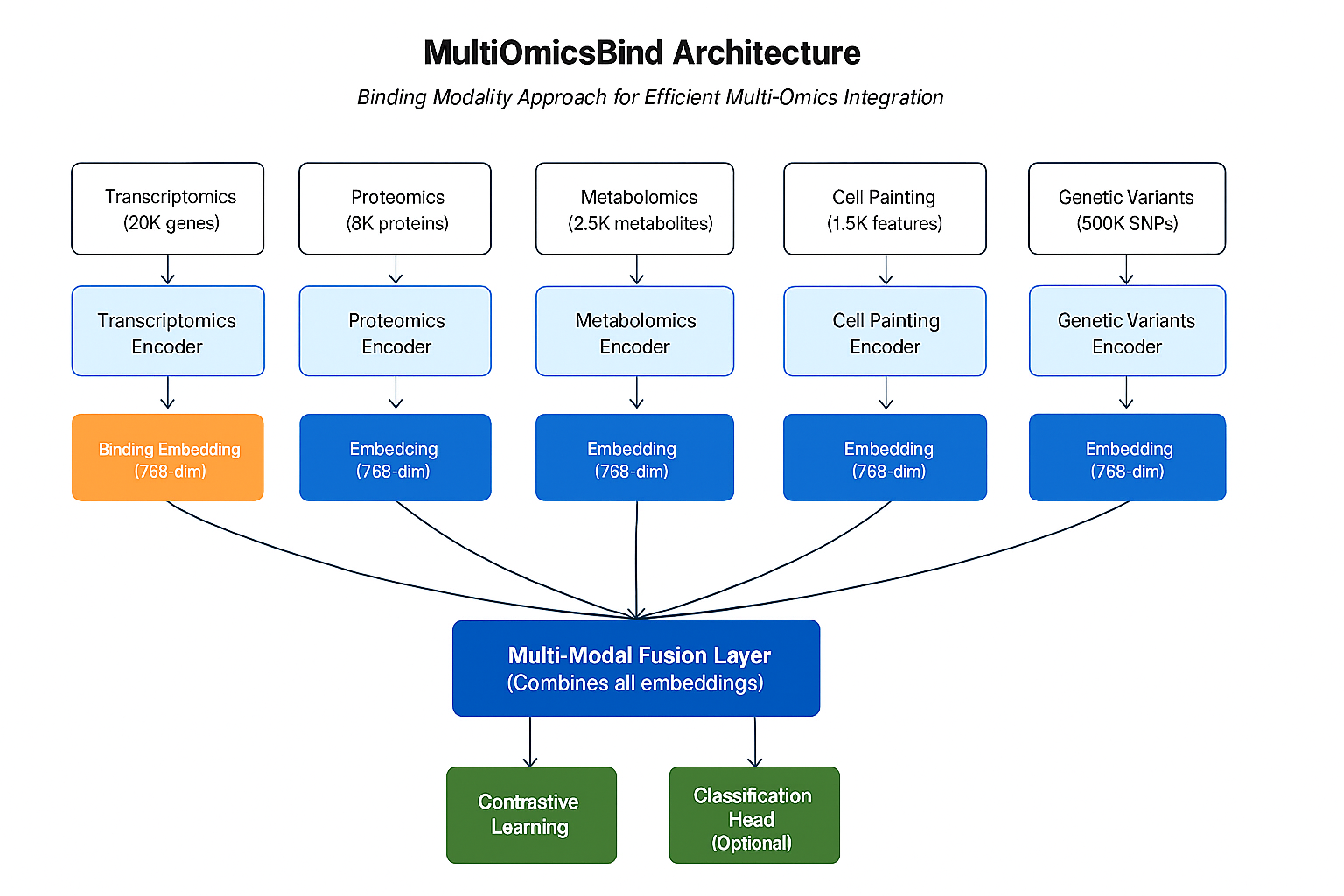
MultiOmicsBind uses a novel approach combining:
- 🎯 Modality-Specific Encoders: Each omics type gets its own specialized neural network encoder
- 🤝 Cross-Modal Alignment: Contrastive learning ensures related samples have similar representations across modalities
- 🎪 Flexible Integration: Learned embeddings can be used for downstream tasks like classification, clustering, or biomarker discovery
The framework learns to create a unified representation space where:
- Samples with similar biological profiles cluster together
- Different modalities of the same sample align closely
- Meaningful biological relationships are preserved
Real-World Applications 🌍
🏥 Precision Medicine
- Integrate genomic, clinical, and imaging data for personalized treatment
- Identify patient subgroups with distinct molecular profiles
💊 Drug Discovery
- Combine compound structure, gene expression, and phenotypic data
- Predict drug responses and identify novel targets
🔬 Biomarker Discovery
- Identify multi-omics signatures for disease diagnosis
- Discover prognostic markers across data types
🧬 Systems Biology
- Understand complex biological networks
- Model multi-scale biological processes
🔗 Get Involved:
⭐ Star the repository on GitHub
🐛 Report bugs and request features
💡 Contribute code and documentation
What are your biggest challenges in multi-omics data integration? Share your thoughts in the comments below! 💭
#MultiOmics #DeepLearning #MachineLearning #Bioinformatics #DataScience #PrecisionMedicine #AI #Research #OpenSource #PyTorch #Genomics #Proteomics #SystemsBiology #LifeSciences #Innovation #BioML #ComputationalBiology
👉 GitHub Repository: MultiOmicsBind