ScAdver: Adversarial Batch Correction for Single-Cell Data
Published:
ScAdver - Adversarial Batch Correction for Single-Cell Data
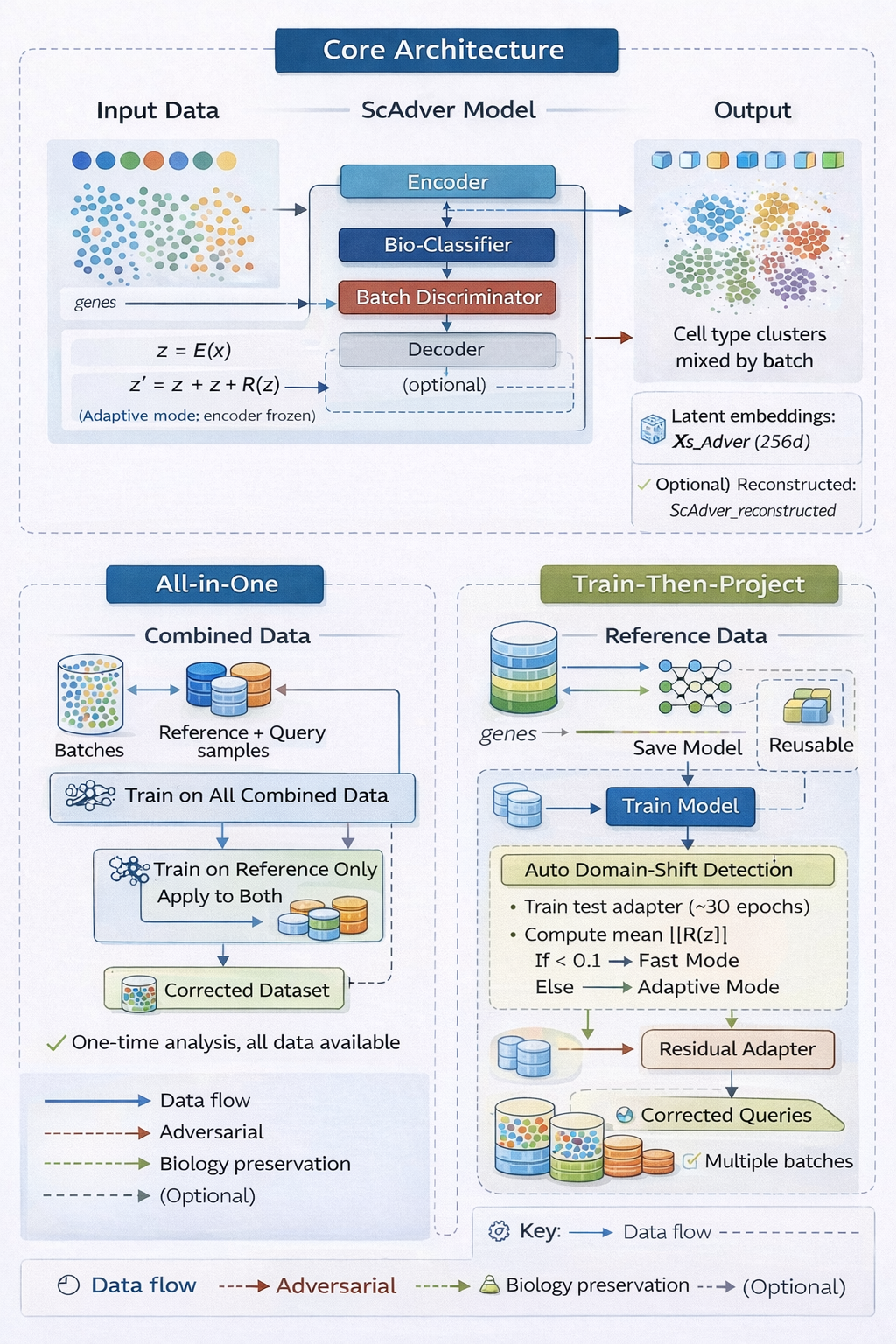
ScAdver removes technical batch effects from single-cell RNA-seq data while preserving biological signal and cell identity. It follows a train-once, project-forever paradigm: train on reference data once, then project unlimited query batches without retraining the core encoder.
Key Features
- Train once, project forever: Reuse a trained encoder across many query batches.
- Fully reproducible:
seed=42triggers internal global seeding in training functions. - Biology preserved: Adversarial training keeps biological structure while removing technical variation.
- Enhanced residual adapter: 3-layer adapter with LayerNorm and GELU for robust domain shifts.
- Distribution alignment: MMD + Moment-Matching + CORAL losses for stable adaptation.
- Probe-gated query projection: Automatic routing in
transform_query_adaptivebased on measured latent shift and overlap. - Fast large-scale mode: Analytical path supports 100k+ cells efficiently.
- Multi-device: CPU, CUDA, and Apple Silicon (MPS).
Installation
pip install git+https://github.com/shivaprasad-patil/ScAdver.git
Workflows
Workflow 1: All-in-One Batch Correction
from scadver import adversarial_batch_correction
adata_corrected, model, metrics = adversarial_batch_correction(
adata=adata,
bio_label='celltype',
batch_label='batch',
epochs=500,
seed=42,
return_reconstructed=True
)
Use this when all data is already available and you want one-time correction.
Workflow 2: Reference-to-Query (Train-Then-Project)
from scadver import adversarial_batch_correction, transform_query_adaptive
# Step 1: train once on reference
adata_ref_corrected, model, metrics = adversarial_batch_correction(
adata=adata_reference,
bio_label='celltype',
batch_label='tech',
epochs=500,
seed=42
)
# Step 2: project query batches as they arrive
adata_query = transform_query_adaptive(
model=model,
adata_query=adata_query,
adata_reference=adata_reference,
bio_label='celltype'
)
Use this when query data arrives over time, comes from a shifted protocol, or you want a reusable deployed model.
Which Workflow to Choose?
| Scenario | Recommended Workflow |
|---|---|
| All data available now, one-time analysis | Workflow 1 |
| Query batches arrive over time | Workflow 2 |
| Large protocol shift (e.g., 10X to Smart-seq2) | Workflow 2 |
| Deploying as a reusable service | Workflow 2 |
How It Works
The encoder is trained adversarially to:
- Keep biological patterns via a bio-classifier.
- Remove batch patterns via an adversarial discriminator.
For query projection, transform_query_adaptive measures latent shift and routes to one of three paths:
- Neighborhood residual path for strong overlap and moderate class count.
- Neural adapter path for moderate-to-large biological diversity (up to 100 classes).
- Analytical mean-shift path for very large class spaces (100+ classes).
Output
- Latent embeddings:
adata.obsm['X_ScAdver'](256-dimensional, batch-corrected). - Reconstructed expression:
adata.layers['ScAdver_reconstructed'](optional, setreturn_reconstructed=True). - Metrics: Biology preservation, batch correction, overall score.
Documentation
- ENCODER_MECHANISM_EXPLAINED.md - Encoder training and projection mechanics.
- RESIDUAL_ADAPTER.md - Residual adapter and adaptation details.
- Pancreas notebook - End-to-end walkthrough.
- PBMC notebook - v2/v3 protocol integration.
GitHub Repository: ScAdver